Prediction driven design of hERG liability free compounds
Optimizing pharmacokinetics and toxicity attributes are profound objectives across all drug discovery projects. The hERG potassium ion channel contributes to the normal electrical activity in the heart. Interference with xenobiotics can result in long QT syndrome and life-threatening arrhythmias. Therefore early detection of the hERG inhibition during virtual screening or medicinal chemistry optimization has essential importance.
The new ChemAxon ADMET plugin group exploits the power of machine learning methods on curated data sets to support the drug design and medicinal chemistry optimization with reliable models and predictions. The first available pre-trained model is targeting the elimination of cardiotoxicity risk during drug discovery by predicting hERG inhibition. We present the model development and assessment data on our hERG pAct regression model. We also discuss the requirements of modern applications interfacing with machine learning models that lead to higher detail prediction results, such as estimated error, applicability domain, and the visualization of similar training compounds.
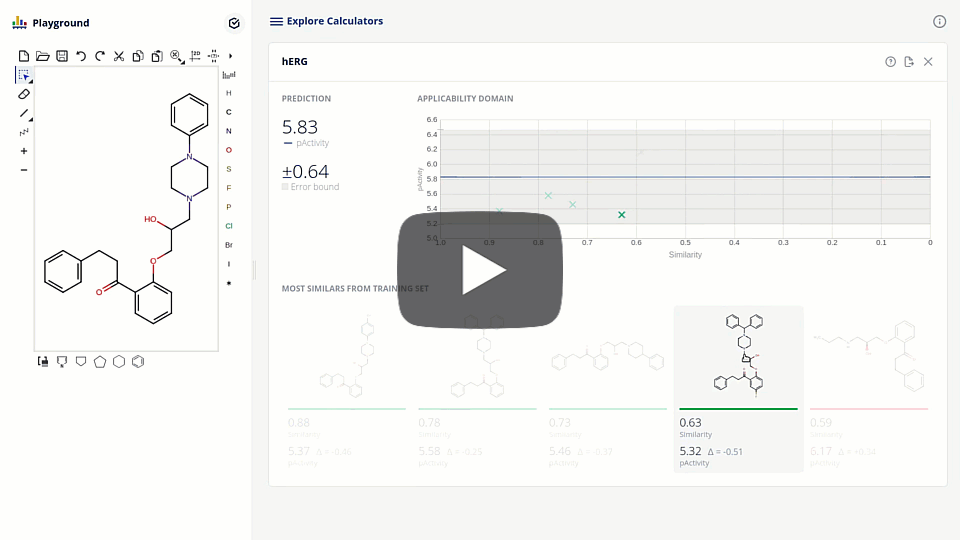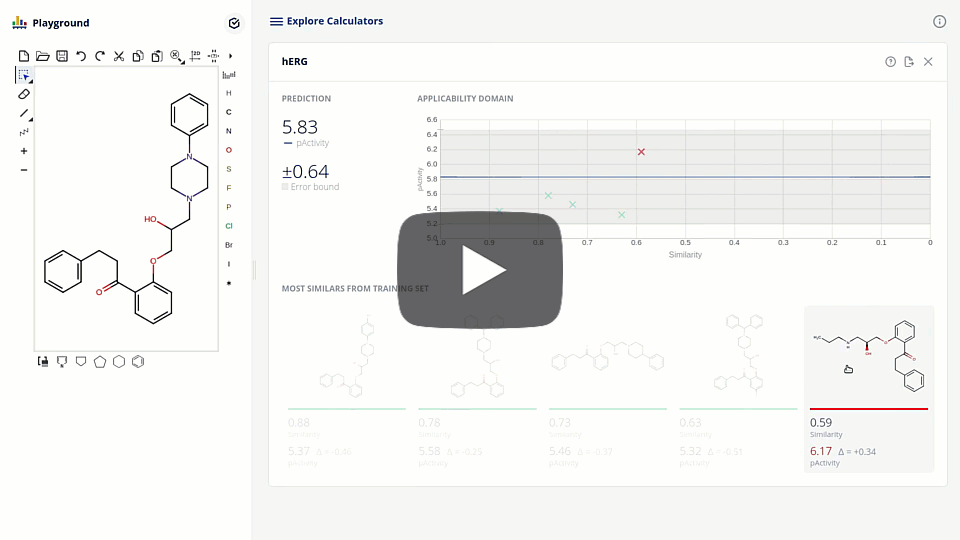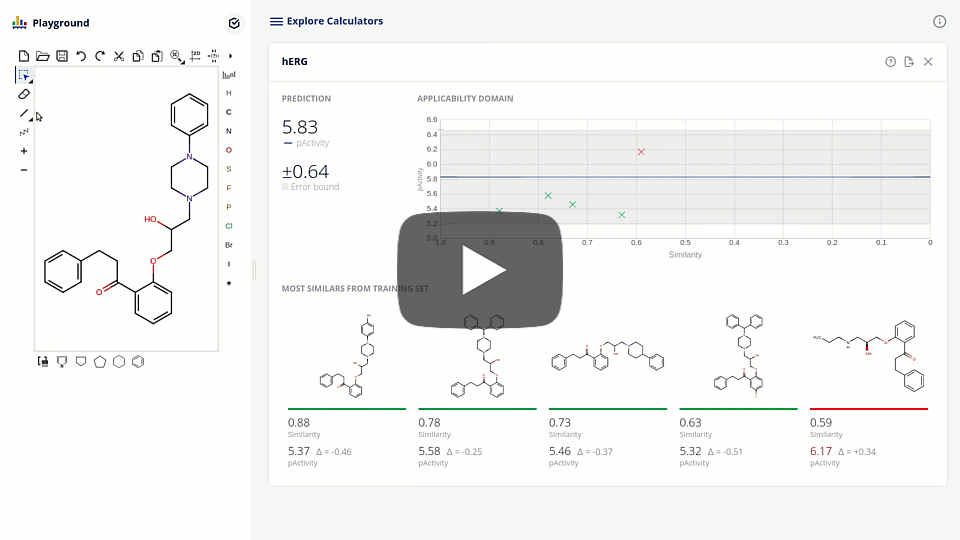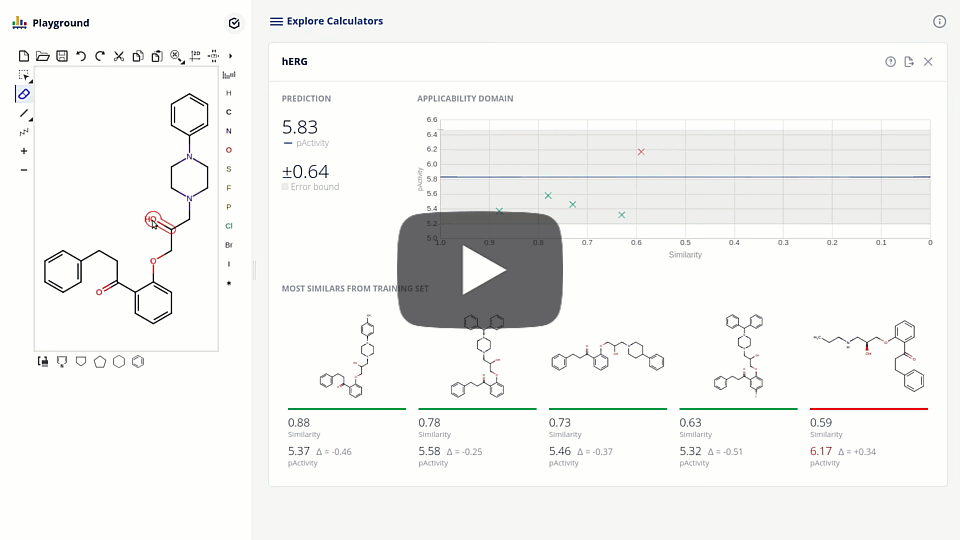
Due to the challenge of optimizing the ADMET parameters and interaction with one single target out of the ~20k proteins present in the human body, drug design is inherently a multi-dimensional task. The ultimate goal is to balance the attributes of the structures to meet the desired target-product profile. Accordingly, design tools are to be organized into smoothly connected systems, where all the necessary information is effectively collected.
We present here the integration of the hERG model into a platform that connects enabling tools, including database searches, phys-chem predictions, computational models with project pipeline. The tool that fosters collaboration, sharing, triaging and prioritizing compound sets in the pre-synthesis phase.